BIOIMAGING 2015
4TH INTERNATIONAL SYMPOSIUM IN APPLIED BIOIMAGING
THE PRE-CLINICAL CHALLENGE IN 3D
CHALLENGE
WELCOME
We are excited to announce that this year we are including a grand Challenge in our symposium.
Besides a prize money of 512 EUR, the winner team will receive one registration waiver and will be invited to present their work orally at a dedicated programme slot in the symposium.
We all know that great advances in science can be achieved with a proper blend between cooperation and competition. The Challenge we propose tries to gain from this perspective and consists of a non-trivial problem in bioimaging, with very high scientific interest.
DESCRIPTION OF THE CHALLENGE
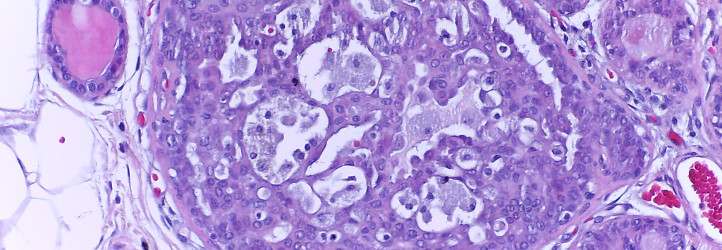
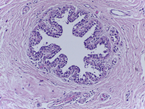
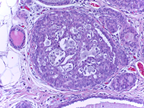
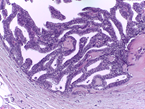
This Challenge is presented in the context of breast cancer and has the goal of pushing forward the methods for automatic classification of tissue malignancy in the field of digital pathology. Tumor classification requires careful examination of hematoxylin and eosin (HE) stained biopsy samples performed by pathologists. Given an image of an HE stain biopsy tissue sample (examples below), the goal is to automatically classify the sample as: a) normal tissue; b) benign lesion; c) carcinoma in situ; or d) invasive carcinoma.
 |
 |
 |
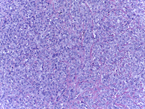 |
| a) normal tissue | b) benign lesion | c) carcinoma in situ | d) invasive carcinoma |
DESCRIPTION OF THE DATASET
A dataset with 140 high-resolution (2048x1536), annotated HE stain images has been produced specifically for this Challenge. The images were all digitized with the same acquisition conditions, and with a magnification of 200x. The dataset has been assembled and annotated/classified by two pathologists of the Diagnostics Unit of Ipatimup - Institute of Molecular Pathology and Immunology at the University of Porto: António Polónia (MD) and Catarina Eloy (MD, PhD).
The dataset contains 30 images from different biopsy samples, for each one of the four classes (normal tissue, benign lesion, carcinoma in situ and invasive carcinoma). These 120 classified images are provided to the Challenge participants as a training dataset. To download the dataset you must first register a participating team (individual submissions are also allowed).
In addition, the dataset contains 20 images, not available to the Challenge participants, for testing and scoring the submitted algorithms (test dataset).
HOW TO PARTICIPATE
In order for individuals or teams to participate in the Challenge they need to register using the: CHALLENGE PARTICIPATION FORM >> .
Once registered, a link will be provided for downloading the training dataset. Use of this dataset for other purposes than the Bioimaging 2015 Challenge is not permitted (please check the Challenge Rules).
Each team needs to submit, until October 16:
- a two pages report/short-paper describing the algorithm, and - an executable code/program with the algorithm for scoring by the Jury; this should be a console application (no GUI) which receives as input an image filename and outputs the classification (normal tissue, benign lesion, carcinoma in situ or invasive carcinoma)
The teams are totally free to choose the methods/techniques to use in the Challenge.
The algorithms can be coded in any language (C, C++, Python, MATLAB, etc.), for either MSWin or GNU-Linux OS, as long as it is possible for the Jury to independently run the code/program (input image; output classification).
Each team can only submit one algorithm.
IMPORTANT DATES
Challenge short-paper and algorithm submission: October 16, 2015
Challenge score results and winner announcement: October 23, 2015
RUA DO CAMPO ALEGRE, 823 | 4150-180 PORTO - PORTUGAL | TEL +351 226 074 900 | EMAIL: BIOIMAGING2015@INEB.UP.PT
| ORGANIZED BY: | CO-ORGANIZED BY: | SUPPORT: | |||||
 |
 |
 |
 |
||||